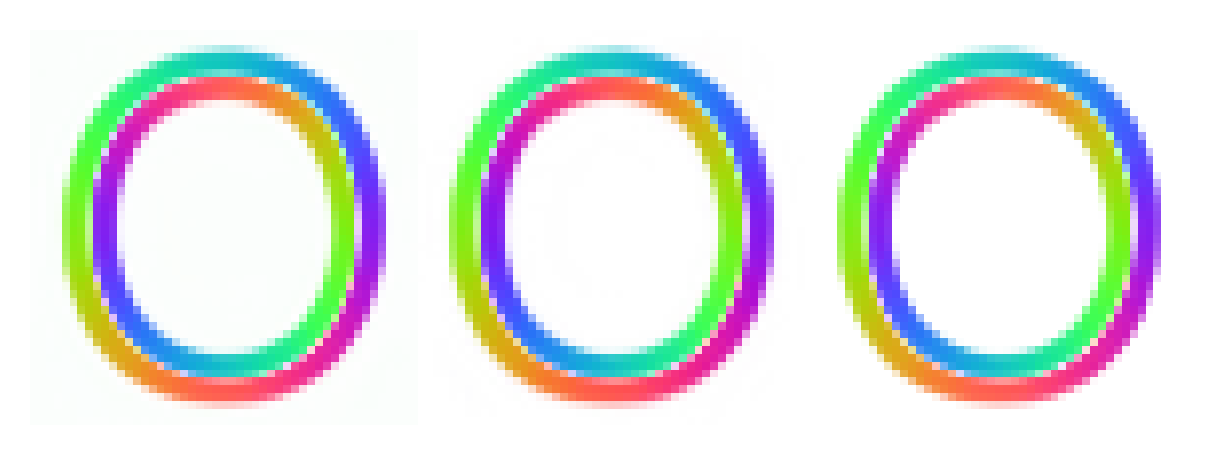Logo#
This is a little cute: Omnipose can "segment" text using the bact_phase_omni model. Semantic segmentation of uniform,
disjoint shapes on a uniform background is absolutely no feat, but it is amusing that a neural network trained purely on
phase contrast images of bacteria gives such reasonable output on something so different from the training set.
Also, the over-segmentation at cusps hints that the network has learned to pick up on local morphology.
To make the Omnipose logo/title/favicon, I first generate some rasterized text images with roughly the same mean diameter as the bacteria in my training set:
Show code cell source
1# Make some text images
2
3from PIL import Image, ImageDraw, ImageFont
4import numpy as np
5import matplotlib.pyplot as plt
6plt.style.use('dark_background')
7import matplotlib as mpl
8%matplotlib inline
9mpl.rcParams['figure.dpi'] = 300
10
11from omnipose.utils import bbox_to_slice
12
13tsizes = [60]
14texts = ["Omnipose","O"]
15imgs = []
16for textsize in tsizes:
17 fonts = [ImageFont.truetype(f, textsize) for f in ["SFNSRounded.ttf"]]
18 # fonts = [ImageFont.truetype(f, textsize) for f in ["Arial.ttf"]]
19 for text in texts:
20 for font in fonts:
21 size = np.array([textsize*len(text)*2, textsize*2])
22 im = Image.new("RGB", tuple(size), "white")
23 d = ImageDraw.Draw(im)
24 center = size/2
25 anchor = "mm"
26 d.text(center, text, fill="black", anchor=anchor, font=font)
27 bbox = d.textbbox(center, text, anchor=anchor, font=font)
28 bbox = [bbox[1],bbox[0],bbox[3],bbox[2]] # reverse x, y
29 im = np.array(im)
30 shape = im.shape[:2]
31 slc = bbox_to_slice(bbox,shape,pad = 3)
32 im = im[slc]
33 imgs.append(im)
34
35
36 fig = plt.figure(figsize=(1,1))
37 fig.patch.set_facecolor([0]*4)
38
39
40 plt.imshow(im)
41 plt.axis('off')
42 plt.show()


Segmentation#
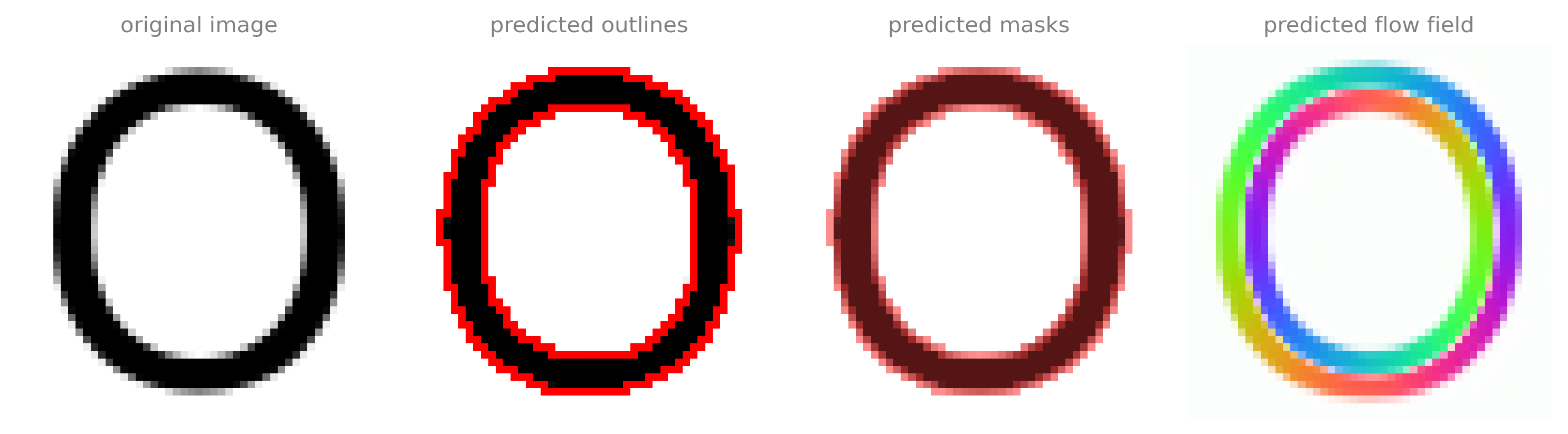
I will then segment these image with the standard settings:
Show code cell source
1from cellpose_omni import plot, models, core
2import omnipose
3
4model_name = 'bact_phase_omni'
5use_GPU = core.use_gpu()
6model = models.CellposeModel(gpu=use_GPU, model_type=model_name)
7
8
9chans = [0,0] #this means segment based on first channel, no second channel
10nimg = len(imgs)
11n = range(nimg)
12
13# define parameters
14mask_threshold = 1
15verbose = 0
16use_gpu = use_GPU
17transparency = True
18rescale=None
19omni = True
20flow_threshold = 0
21resample = True
22cluster = False
23
24masks, flows, styles = model.eval([imgs[i] for i in n],
25 channels=chans,
26 rescale=rescale,
27 mask_threshold=mask_threshold,
28 transparency=transparency,
29 flow_threshold=flow_threshold,
30 omni=omni,resample=resample,
31 verbose=verbose,
32 cluster=cluster)
33
34
35mpl.rcParams['figure.dpi'] = 300
36plt.style.use('dark_background')
37
38for idx,i in enumerate(n):
39
40 maski = masks[idx] # get masks
41 bdi = flows[idx][-1] # get boundaries
42 flowi = flows[idx][0] # get RGB flows
43
44 # set up the output figure to better match the resolution of the images
45 # f = 10
46 # szX = maski.shape[-1]/mpl.rcParams['figure.dpi']*f
47 # szY = maski.shape[-2]/mpl.rcParams['figure.dpi']*f
48 szX,szY = 10,10
49 fig = plt.figure(figsize=(szY,szX*4))
50 fig.patch.set_facecolor([0]*4)
51
52 plot.show_segmentation(fig, omnipose.utils.normalize99(imgs[i]),
53 maski, flowi, bdi, channels=chans, omni=True, interpolation=None)
54
55 plt.tight_layout()
56 plt.show()
I landed on this font because it is one of Apple's system defaults (and therefore works well with the system fonts used on our website when viewed on Apple devices), and I chose this scale because it is very close to bacteria and showed a good amount of 'segmentation' in the M, N, P, and E from purely local morphology (cusps). This gives reasonable output at higher-resolution text (wider 'cells'), but it starts to hallucinate output between objects if the size gets too large.
Adjusting transparency#
The transparency (alpha channel) is set by the flow magnitude, and the color (RGB channels) is set by the flow angle according to a shifted sinebow relation:
angles = np.arctan2(dP[1], dP[0])+np.pi
r = ((np.cos(angles)+1)/2)
g = ((np.cos(angles+2*np.pi/3)+1)/2)
b =((np.cos(angles+4*np.pi/3)+1)/2)
(a is just a constant, not alpha). The slight tinge of green comes from the fact that np.arctan2(0,0)=0 and ((np.cos(0+2*np.pi/3)+1)/2) = 1/4. I'll try two ways to remove it:
first by removing any average background bias, second by adjusting the alpha channel so that the background alpha is 0 on average.
Show code cell source
1import skimage.io
2import os
3from omnipose.utils import normalize99, rescale
4from scipy.ndimage import zoom
5from pathlib import Path
6
7omnidir = Path(omnipose.__file__).parent.parent
8basedir = os.path.join(omnidir,'docs','_static')
9names = ['logo.png','icon.ico']
10ext = '.png'
11
12for idx,i in enumerate(n):
13
14 maski = masks[idx]
15 flowi = flows[idx][0]
16 dPi = flows[idx][1]
17 bias = [np.mean(d[maski==0]) for d in dPi]
18 angle = np.arctan2(bias[1], bias[0]) / np.pi
19 print('avarage bias is {}, average angle is {} pi rad.'.format(bias,angle))
20 dPi_new = np.stack([np.clip(d - b,-np.inf,np.inf) for d,b in zip(dPi,bias)])
21 flowi_new = plot.dx_to_circ(dPi_new,transparency=True)
22
23 flowi_3 = flowi.copy()
24 alpha = flowi_3[...,-1]
25 flowi_3[...,-1] = rescale(np.clip(alpha-np.mean(alpha[maski==0]),0,np.inf))*255
26
27 f = 30
28 szX = maski.shape[-1]/mpl.rcParams['figure.dpi']*f
29 szY = maski.shape[-2]/mpl.rcParams['figure.dpi']*f
30 fig = plt.figure(figsize=(szY,szX*4))
31 fig.patch.set_facecolor([0]*4)
32
33 plt.imshow(np.hstack([flowi,flowi_new,flowi_3]))
34 plt.axis('off')
35 plt.show()
36 # aplha channel correction is the winner
37 # also rescale the image without interpolation so that, when displayed as favicon etc., it is not as smoothed out - we want to show real output
38 skimage.io.imsave(os.path.join(basedir,names[idx]),zoom(flowi_3,(3,)*(flowi.ndim-1)+(1,),order=0))
39
Turns out that subtracting off the flow component bias introduces some over-correction in places, leading to some discoloration. So, alpha adjustment it is. It might be hard for you to see it, but I can. This level of pixel-peeping is how I made my ground-truth data ;-)
Exporting#
Favicons need to be a particular resolution. For now I am making a multi-scale .ico, but that isn't working properly on Safari (too pixelated). Seems like multiple separate PNGs is the way to go moving forward.
1from PIL import Image
2filename = os.path.join(basedir,names[-1])
3zimgs = []
4
5for j,sz in enumerate([(32,32), (128,128), (180,180), (192,192)]):
6 scale = np.array(sz)/np.array(flowi_3.shape[0:2])
7 zimg = zoom(flowi_3,tuple(scale)+(1,),order=(np.max(scale)<1))
8 zimgs.append(zimg)
9 # plt.imshow(zimg)
10 # plt.axis('off')
11 # plt.show()
12 # zimg.shape
13
14icon = Image.fromarray(zimgs[0], 'RGBA')
15icon.save(filename,append_images=[Image.fromarray(z, 'RGBA') for z in zimgs[1:]])